Data Management
Resources for developing DMPs and engaging with KBase.
Current language will be adjusted as these plans and guidance evolve.
Data Management Plans (DMPs)
Required for all publishable data, metadata, and software from research funded by the the Genomic Science Program (https://genomicscience.energy.gov/datasharing/) and Department of Energy (https://www.energy.gov/datamanagement/doe-policy-digital-research-data-management).
Data Management Plans, or DMPs, should address how samples, data, software, and any other research products generated by the project will be accessed, shared, and preserved, and include ownership/responsibility considerations. These topics support the generation of Findable, Accessible, Interoperable, and Reusable (FAIR, Wilkinson et al. 2016) scientific research products. Quality metadata about the research products, especially samples, data, and the provenance on how each were generated, also support comparability and reproducibility. We recommend reviewing the 10 Simple Rules and Getting and Giving Credit for Data (Wood-Charlson et al. 2022) for additional details and resources.
For each topic, we have provided information and example text on how KBase adheres to community standards, enables sharing and public access, and fits into the broader architecture of DOE-funded resources that support a complete data lifecycle (Figure 1).
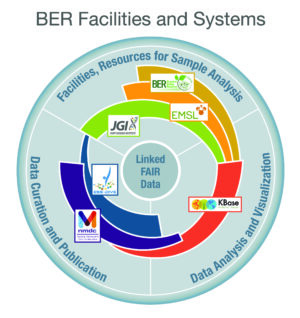 Fig. 1 Data and Analysis Ecosystem BER researchers and facilities produce a wealth of data for predictively understanding complex biological systems. To fully leverage this data, BSSD supports integrative computational and data science platforms to facilitate community access, analysis, and sharing of omics, imaging, and other data types. [Courtesy Lawrence Berkeley National Laboratory] (U.S. DOE. 2021. Biological Systems Science Division Strategic Plan, DOE/SC-0205. U.S. Department of Energy Office of Science (genomicscience.energy.gov/2021bssdstrategicplan/).
Fig. 1 Data and Analysis Ecosystem BER researchers and facilities produce a wealth of data for predictively understanding complex biological systems. To fully leverage this data, BSSD supports integrative computational and data science platforms to facilitate community access, analysis, and sharing of omics, imaging, and other data types. [Courtesy Lawrence Berkeley National Laboratory] (U.S. DOE. 2021. Biological Systems Science Division Strategic Plan, DOE/SC-0205. U.S. Department of Energy Office of Science (genomicscience.energy.gov/2021bssdstrategicplan/).
Samples and data
Describe the types of samples that will be generated; the protocols, tools, and instruments used to generate data; and any analysis the data will undergo. Include descriptions (size, format, frequency, etc) of any existing or planned datasets.
Samples
While not a sample repository, KBase does support community standards used to describe sample metadata. Examples include: International Geo/General Sample Number (IGSN) – a community-standard persistent identifier for physical samples, and NCBI Biosamples (see documentation on KBase samples for more information). For microbiome-specific samples, we recommend also reviewing the National Microbiome Data Collaborative’s DMP page on “Microbiome Community Standards & Repositories.”
Example text: KBase supports samples by storing sample metadata that adheres to community standards (or robust user-defined descriptors of samples where standards have not yet been established) and making it available for comparative analyses and exploration in the KBase platform.
Data
KBase supports a wide range of data types and file formats related to systems biology research, including genomes, annotations, metagenomes, expression, protein-protein interactions, models of organismal and community metabolism, gene regulation, and sample metadata. See links for a complete list of KBase data types and file formats and KBase Data Sources.
Example text: All curated data can be uploaded (and downloaded) using standardized file formats. Data will be integrated into KBase for open-access data sharing, and reproducible analysis through Narratives. Any Narrative used to share results and/or generated for publication will be assigned a DOI and cited in the publication’s reference section. All data contributed to KBase are immediately tracked for reuse within the KBase provenance architecture. KBase provides views and copies/download statistics for all Narratives with a DOI, which demonstrates the broader impact of the data through its use by others.
Software
KBase supports open source software and best practices as a resource developed to provide public access to cutting-edge software tools relevant to the mission of the Department of Energy Office of Science. It also provides a means for users to share their data and optimized software workflows. All KBase code is available in the KBase Github repository, as a community-standard, open source platform for making software and code widely available.
Example text: All software tools described are open source and will be freely available through KBase and Github, including the KBase GitHub repository. Documentation will be provided to users to empower adoption for their research studies. All software housed within KBase is developed under open source licenses (e.g., the MIT Open Source License). Impact of developer contributions are tracked as the number of Narratives a tool has been used in and how many times it has been run. In addition, all KBase Apps are cited within the Narrative and automatically linked to any downstream Narrative DOIs generated when a user is publishing their research.
Access, sharing, and preservation
KBase is free and open to anyone able to access a web browser and create an account. Data and analysis workflows can be made accessible outside the KBase login by creating a “static” HTML snapshot of the Narrative. Full access (e.g., add data, run tools, or download) requires viewers to create an account, as that supports provenance and reuse tracking.Therefore, it is recommended to provide sufficient documentation in the Narrative such that readers can explore your results prior to being asked to login.
A specific version of a KBase Narrative can be saved and assigned a DOI, by request. The Narrative DOI. The default reuse policy for Narratives with a DOI is a Creative Commons CC0 license, allowing unlimited reuse. Narrative authors may request a CC-BY-4.0 license (reuse with attribution) for a Narrative as an analysis workflow, but all data within the Narrative remain CC0.
All KBase user data are backed up to the Amazon S3 cloud services, ensuring that the full system could be restored if necessary. If KBase were to no longer be supported, KBase has a partnership with the California Digital Library to migrate all data associated with a Narrative with a DOI to Dryad for long-term access and storage.
Example text: Access, sharing, and preservation of data will be maintained through KBase’s existing functionality of daily backups to Google S3. By request, datasets can be assigned a DOI with a CC0 license. Relevant metadata will be included in the DOI landing page and full accessibility will be available within the KBase Narrative environment. As necessary, KBase will preserve all data and metadata associated with a DOI in a repository affiliated with the California Digital Library for long-term access and storage.
Ownership and responsibilities
The DMP should ensure that roles and responsibilities are accounted for and it is clear how the requirements for data management will be met. It should also be clear how responsibilities will be transferred if personnel leave the project. In KBase, the creator of a Narrative or Organization is permanently assigned. However, full administrative privileges can be granted (and revoked for) to other KBase users.
Example text: Primary ownership of data and organizations in KBase will be coordinated by (name/position). In the event (name/position) is no longer on the project, the administration of these resources will pass to (second name/position).
KBase also supports bioinformaticians to contribute code and analysis tools to the KBase open source project. Community developers are trained in the KBase software development kit (SDK) and supported throughout the KBase software release process. It is expected that community developers maintain and update their code as necessary.
Example text: KBase supports community developers to incorporate their software into the KBase open source project. By doing so, community developers agree to follow best practices, and maintain and update their code as necessary.
How to cite KBase: Arkin AP, Cottingham RW, Henry CS, Harris NL, Stevens RL, Maslov S, et al. KBase: The United States Department of Energy Systems Biology Knowledgebase. Nature Biotechnology. 2018; 36: 566. doi: 10.1038/nbt.4163
How to cite the KBase DMP: Arkin AP, et al. DOE Systems Biology Knowledgebase (KBase) Data Management Plan. DMPTool. 2024. doi: 10.48321/D13BA96b4d
Resources on engaging with KBase
Opportunities through the Genomic Science Program Science Focus Areas and Funding Opportunity Announcement solicitations and invitations for research funding from the Department of Energy Office of Science. The following programs are available focused on workforce development and skill-building opportunities.
DOE-sponsored and collaborating National Lab programs:
- Department of Energy’s Office of Science (DOE-SC) Workforce Development for Teachers and Scientists (WDTS) programs (https://science.osti.gov/wdts)
- LBNL
- Workforce Development and Education (https://education.lbl.gov/internships/)
- K12 STEM Education and Outreach (https://k12education.lbl.gov/)
- ORNL: Educational Programs (https://education.ornl.gov/)
- ANL: Educational Programs and Outreach (https://www.anl.gov/education)
- BNL: Workforce Development and Science Education (https://www.bnl.gov/education/)
KBase supported and affiliated programs:
- KBase Educators supports and collaborates with educators and students at approximately 162 in the US and in 27 countries across 6 continents.
- Regularly hosts summer high school, undergraduate, and graduate interns.
- Involved with the NSF Research Experience for Undergraduates National Summer Undergraduate Research Project (NSURP; https://nsurp.org/).
Example text: KBase supports workforce development toward skilled researchers. The KBase Educators program supports and collaborates with educators and students at approximately 160 instutitions in the US and in 27 countries across 6 continents. The team regularly hosts high school, undergraduate, and graduate interns. KBase is also involved with the NSF Research Experience for Undergraduates National Summer Undergraduate Research Project (NSURP; https://nsurp.org/), a remote program built to support STEM college and university students.
