
A knowledgebase for predictive biology
KBase enables users to analyze, share, and collaborate using data and tools designed to help build increasingly realistic models for biological function.


What is KBase?
The Department of Energy Systems Biology Knowledgebase (KBase) is a knowledge creation and discovery environment designed for biologists and bioinformaticians. KBase integrates a variety of data and analysis tools, from DOE and other public resources, into an easy-to-use platform that leverages scalable computing infrastructure to perform sophisticated systems biology analyses. The platform is a freely available and developer extensible platform where scientists can analyze their own data within the context of public data, using a variety of open-source bioinformatics apps to power their workflows. The collaborative Narrative interface provides a digital notebook the allows users to research together and publish their analysis, data, and code with a persistent link and DOI identifiers to support reproducibility.
KBase is funded by the DOE Biological and Environmental Research (BER) program. DOE BER sponsors user facilities and resources such as the Joint Genome Institute (JGI), the Environmental Molecular Sciences Laboratory (EMSL), the National Microbiome Data Collaborative (NMDC), and Environmental System Science Data Infrastructure for a Virtual Ecosystem (ESS-DIVE).
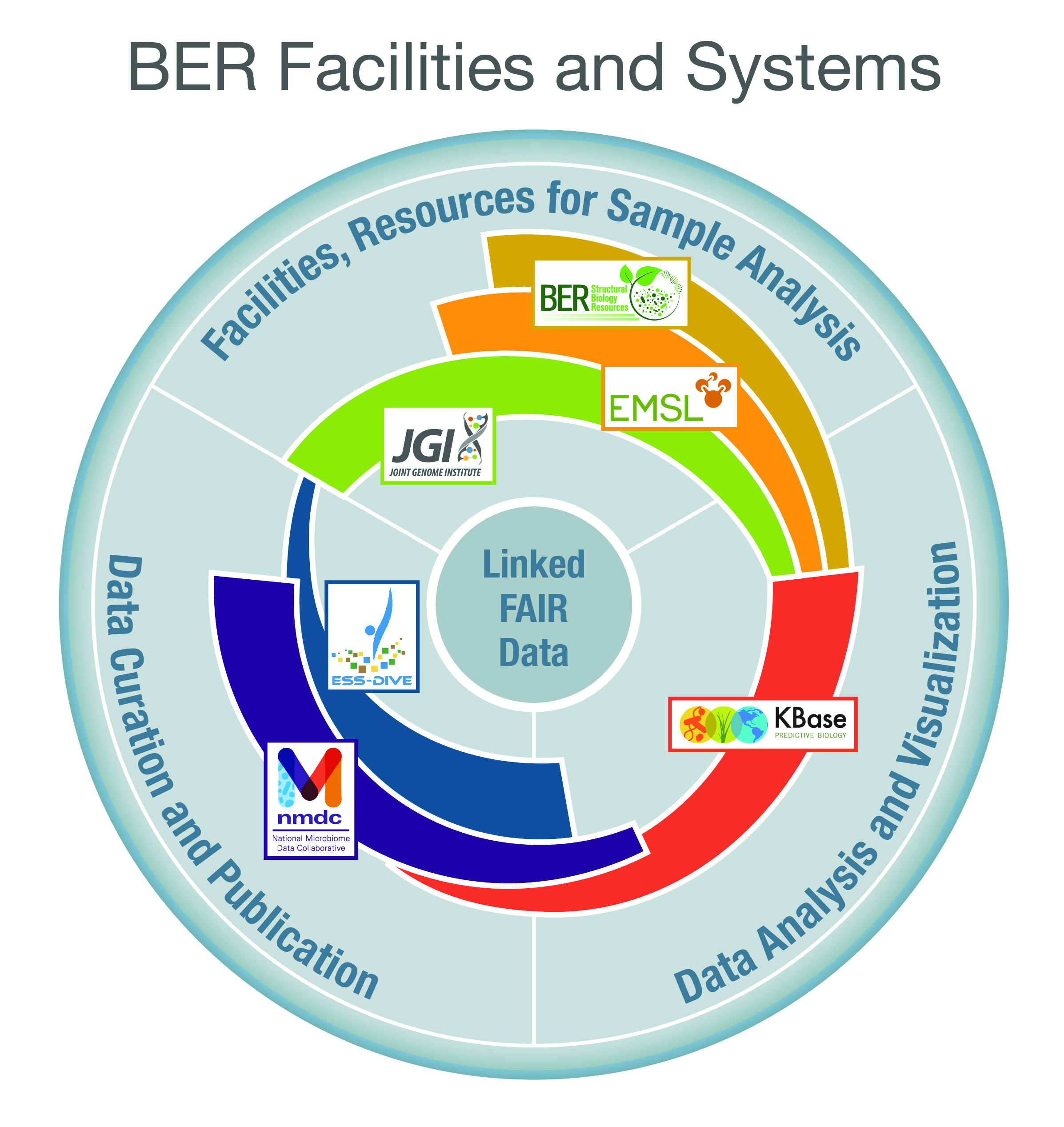
KBase integrates a variety of data from the DOE and other public services into a data model designed around findable, accessible, interoperable, and reusable (FAIR) principles. KBase enables users to analyze their data in the context of public data and share their findings with digital object identifiers (DOIs).
KBase enables bioinformatics developers to add open-source analysis tools that run on KBase’s computational architecture and are available to all users. Users can perform analyses that combine multiple types of ‘omics and biological data to investigate organisms and their communities.
KBase supports sharing of data by researchers and institutions, facilitating collaboration to accelerate science. Our mission is to build a knowledgebase for systems biology, where researchers share discoveries, publish reproducible workflows, and get credit for their work.

Read about KBase in Nature Biotechnology
For a comprehensive overview of KBase and its scientific impact, this publication details the unique features and infrastructure of the KBase platform through several scientific use cases.
Meet the KBase Team
KBase is run by an interdisciplinary and collaborative team led by Lawrence Berkeley National Laboratory with participation from Argonne, Brookhaven, and Oak Ridge National Laboratories.
Also involved in the multi-institutional program are Cold Spring Harbor Laboratory, the University of Illinois at Urbana-Champaign, and the University of Tennessee.
Our key external partners are DOE’s Joint Genome Institute, Environmental Molecular Sciences Laboratory, Bioenergy Research Centers, and several of the Genomic Science Program’s Scientific Focus Areas (SFAs). Several university projects are also important contributors.
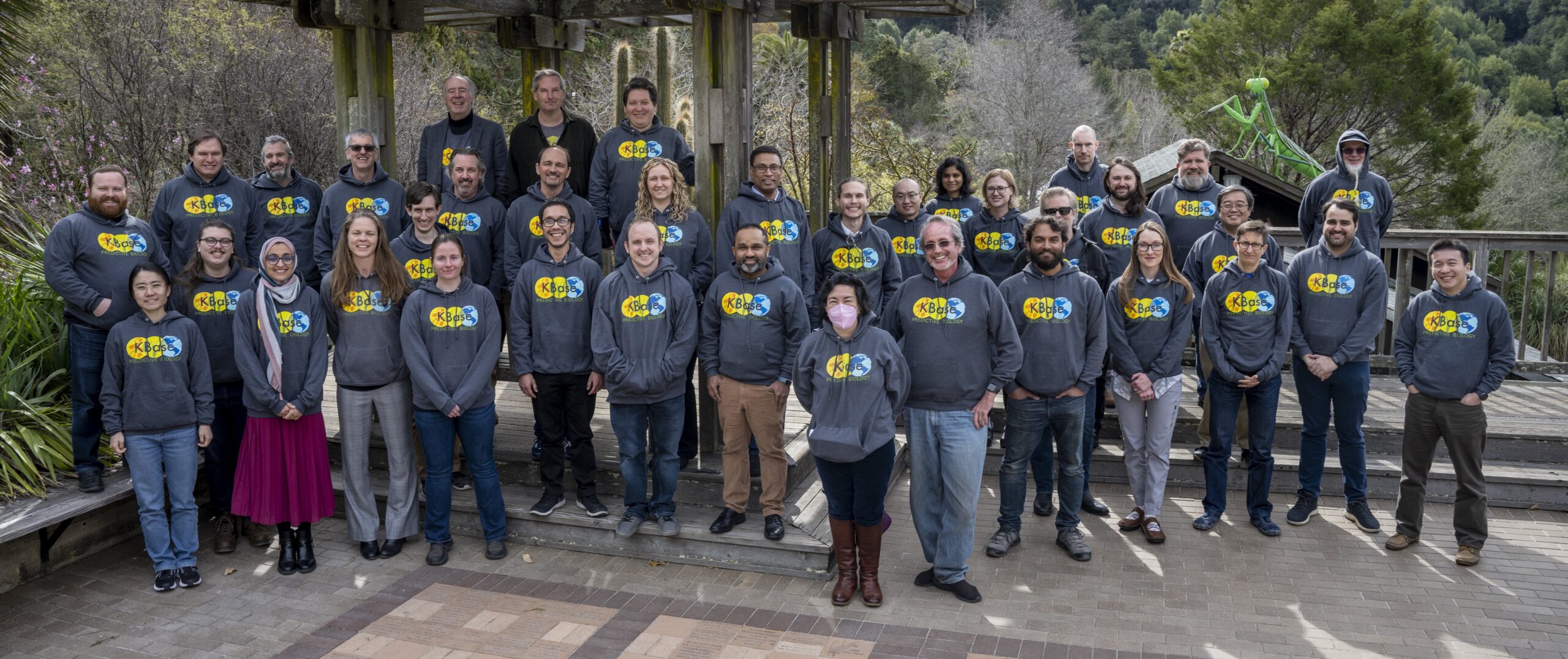 Members of the DOE Energy Systems Biology Knowledgebase (KBase) hold their annual All-Hands meeting where the group discusses past successes, issues that need to be resolved, and plan for future projects that they will undertake, at the UC Berkeley Botanical Gardens, Berkeley, California, 02/22/2023. Photo credit: Thor Swift
Members of the DOE Energy Systems Biology Knowledgebase (KBase) hold their annual All-Hands meeting where the group discusses past successes, issues that need to be resolved, and plan for future projects that they will undertake, at the UC Berkeley Botanical Gardens, Berkeley, California, 02/22/2023. Photo credit: Thor Swift
Funding: KBase is supported as part of the Genomic Sciences Program funded by the U.S. Department of Energy, Office of Science, Office of Biological and Environmental Research (Award Numbers DE-AC02-05CH11231, DE-AC02-06CH11357, DE-AC05-00OR22725, and DE-AC02-98CH10886).
Principal Investigators
Team Leads
Scientific Advisory Committees

Adam is an expert in the comparative systems and synthetic biology of microbes and is dedicated to a model-driven approach to experimental science. He is a senior faculty scientist in the Environmental Genomics and Systems Biology Division at the Lawrence Berkeley National Laboratory and he is the Dean A. Richard Newton Memorial Professor of Bioengineering at the University of California, Berkeley where he has been since 1998. He is Technical Co-Manager of the ENIGMA SFA and directs the Center for Utilization of Biological Engineering in Space. He was one of six recipients of the 2013 Ernest Orlando Lawrence Award, the Department of Energy’s highest scientific honor.

Chris is a scientist at Argonne National Laboratory, a fellow at the University of Chicago, and an adjunct professor at Northwestern University. He is an expert in computational biology with a focus on the prediction of phenotype from genome through the use of comparative genomics, metabolic modeling, and dynamic cellular community models. He received the Jay Bailey Young Investigator Best Paper in Metabolic Engineering Award in 2012.

Bob has extensive experience developing computational and data management tools and systems for genetics, genomics and systems biology research with a background in bioinformatics and management including at the Baylor College of Medicine Human Genome Center as Co-Director of the Informatics Core, Operations Director of the Genome Database at Johns Hopkins University School of Medicine, and Vice President of Computing at Celltech Chiroscience, a UK biopharmaceutical company developing drugs based on gene targets. In 2008 Cottingham moved to Oak Ridge National Laboratory where he is Group Leader for Computational & Predictive Biology.

Elisha M Wood-Charlson is KBase’s User Engagement Lead. She has a PhD and 10+ years of experience as a microbial ecologist focused on host-microbe-virus interactions in the marine environment. Since leaving the research bench, she has moved into the realm of scientific community engagement, with the goal of making microbiome data science more efficient through effective collaboration, building trust in online communities, and developing shared ownership throughout the scientific process.


Gazi Mahmud is seasoned professional with two decades of extensive industry expertise, specializing in the dynamic realms of Big Data and Enterprise Architecture. His core focus lies in seamlessly integrating Data Engineering, Data Science, and modern ML/AI Engineering operations at scale, reflecting his adeptness in orchestrating cross-functional collaborations across diverse domains.
Gazi possesses hands-on experience and visionary insights in design and implementation of data-led organizational transformations, encompassing data architecture modernization, ML/AI integration, and enterprise data governance. Notably, Gazi worked across innovative technology startups and larger high tech companies delivering compelling data narratives that not only foster AI ethics but also champion model explainability within the state of the art Data Science workflow initiatives.
Leading by example, he has navigated expansive projects aimed at crafting innovative data and AI transformation strategies for industry leaders. His expertise extends to implementing continuous delivery capabilities, underscored by meticulous observability, to ensure seamless execution of unified data and analytics platform modernization use cases.

Looking for information on tools and resources?
Check out KBase Documentation for our Getting Started guide and information on tools in the App Catalog. KBase is a fully open source software project available on GitHub.
