
Learn how KBase can help you
Explore tutorials, documentation, and FAQs about the KBase platform.
Get Started with KBase
Watch our quickstart video to learn about the basics of the KBase platform.
Apps for Analysis
KBase apps allow scientists to analyze data with customizable and modular workflows. Apps interoperate seamlessly to enable a range of analysis pathways. The App Catalog lists all of the currently available apps. KBase scientists and our development community build apps using our Software Developer Kit (SDK). The number of apps available in KBase increases as members of the community use our SDK to integrate their analysis tools into the KBase platform.
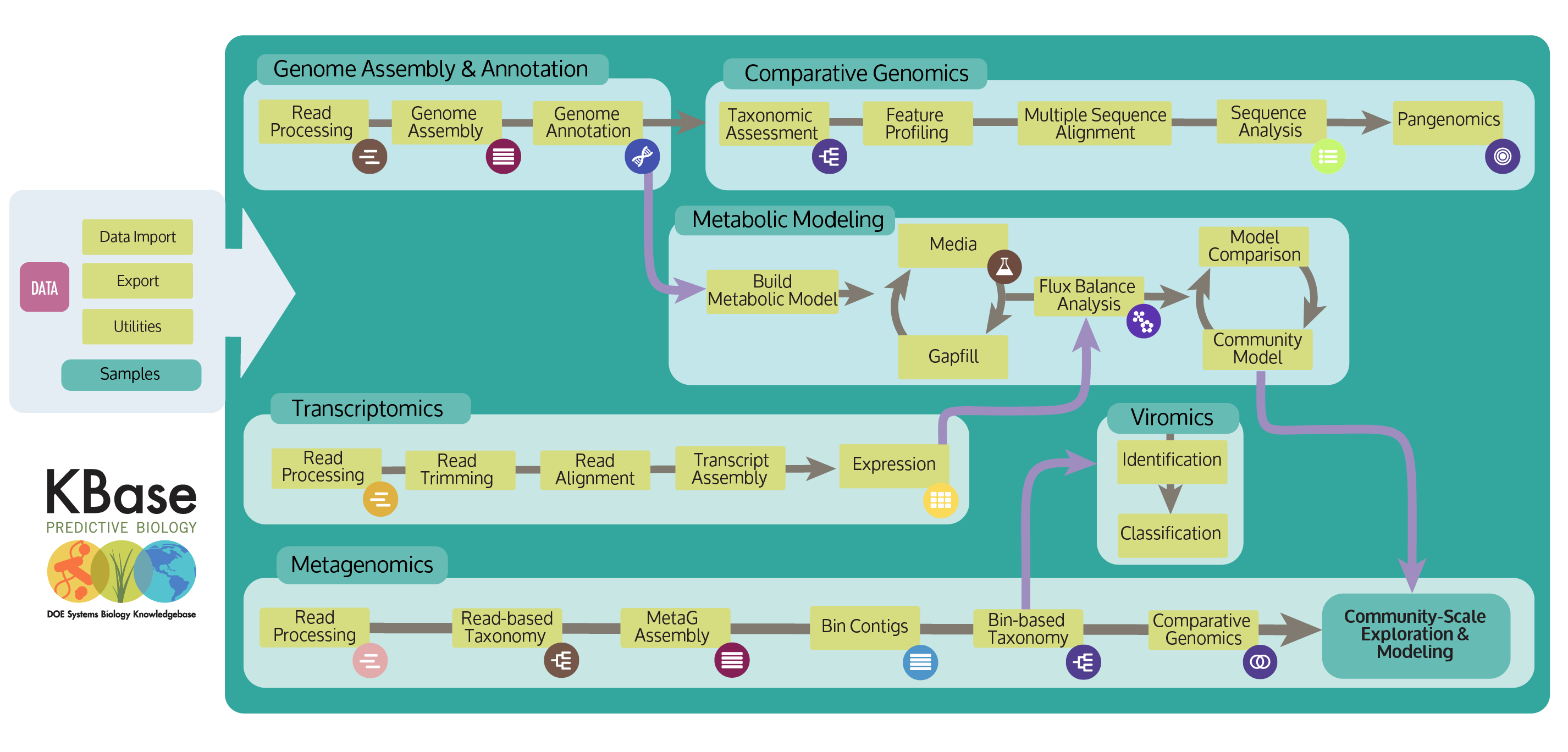
KBase provides multiple Apps for de novo assembly of Next-Generation Sequencing (NGS) reads from various sequencing platforms. These assemblies can be annotated with RAST or Prokka, enabling you to explore structural and functional features.
App ListKBase has a suite of Apps to perform various sequence analyses. They provide the ability to assess quality, search for matches within local databases, and perform multiple sequence alignments.
App ListKBase has a suite of Apps supporting the reconstruction, prediction, and design of metabolic models in microbes and plants. Genome-scale metabolic models can be used to explore an organism’s growth in specific media conditions, determine which biochemical pathways are present, optimize production of an important metabolite, identify high flux pathways, and more.
App ListKBase offers a powerful suite of expression analysis Apps. Starting with short reads, you can use the tool suite to assemble and quantify long transcripts, identify differentially expressed genes, cluster them and analyze them as functionally enriched modules. You can also compare the expression data with the flux when studying metabolic models in KBase and identify pathways where expression and flux agree or conflict.
App ListKBase provides multiple comparative genomics and phylogenetic analysis Apps to enable researchers to understand evolutionary relationships between organisms and explore structural and functional variance across genomes.
App ListKBase Tutorial Narratives
These Narrative tutorials provide step-by-step examples that show how to use KBase tools and data to perform useful analyses. You can copy these Narratives and rerun the steps, modify the analyses, or try on your own data.
Learn how to extract metagenome-assembled genomes (MAGs) from your microbial community sequencing data to obtain high-quality genomes.
View in Nature Protocols.
This Narrative demonstrates processing a single metagenomic dataset from the Global Ocean Viromes dataset, identifying the viral sequences using VirSorter, and subsequently classifying them with vConTACT2.
EMSL Summer School 2021
Learn how to use visualization tools, analysis, and modeling, of multi-omics data for understanding biochemical pathways in this virtual course from EMSL.
Genome Analysis in KBase Series focuses on how to use various tools to analyze genomic sequencing data in KBase. This second Narrative demonstrates a workflow for searching and identifying features within…
Genome Analysis in KBase Series focuses on how to use various tools to analyze genomic sequencing data in KBase. This first Narrative shows how to draft isolate genomes in…
EMSL Summer School 2020
Learn how to incorporate microbial metagenomic and environmental metabolite data from watershed ecosystems into metabolic and community modeling using computational frameworks in this virtual course.
Join the community of educators using KBase to teach bioinformatics and computational biology!
Why KBase?
While KBase is routinely used for advanced analysis by scientists around the world,…
KBase has a suite of tools and data that support the reconstruction, prediction, and design of metabolic networks in microbes. Starting from genome sequence data, this Narrative demonstrates how to use…
PlantSEED was designed to streamline the process of annotating plant genome sequences, constructing metabolic models based on genome annotations, and using models to predict gene functions. This Narrative tutorial demonstrates…
KBase’s RNA-seq pipeline uses Tuxedo tools (including Bowtie, TopHat, Cufflinks, Cuffmerge and Cuffdiff) to quantify gene expression from RNA-seq reads. The resulting expression matrix can be used by other KBase…
Upcoming webinars
Learn about using KBase for your systems biology and bioinformatics research with one of our upcoming webinars. All past webinars are also available for viewing.

Previous webinars
Learn how to import data into KBase, process and assemble short and long sequencing reads, and annotate features on the genome. This session will also how to construct feature sets…
This webinar will introduce the basics of metabolic modeling in KBase. Learn how to construct and gapfill metabolic models, perform flux balance analysis, and explore the metabolic capacity of your…
Learn the basics of KBase and best practices for your research. This webinar introduces the fundamentals of the KBase platform, including data upload, using the Narrative interface, running apps, and…
Kick off the academic year with KBase Educators for an orientation to the Educators Community and teaching resources! We will go over how Narratives and teaching resources are laid out…

Looking for information on tools and resources?
Check out KBase Documentation for our Getting Started guide and information on tools in the App Catalog. KBase is a fully open source software project available on GitHub.
