

Welcome!
The 2020 Multiscale Microbial Dynamics Modeling course was a virtual course hosted by the Environmental Molecular Sciences Laboratory (EMSL) and Pacific Northwest National Laboratory (PNNL) Subsurface Biogeochemical Research group, and featured experts from the Joint Genome Institute and KBase.
Materials from this course cover how to incorporate microbial metagenomic and environmental metabolite data from watershed ecosystems into metabolic and community modeling using computational frameworks, such as KBase and PFLOTRAN. The curriculum includes lectures and software and data analysis tutorials. All of these materials are freely accessible to the community as part of the 2020 Microbial Dynamics Summer School Organization in KBase. Create a KBase account and request to join the KBase Org to get started today!
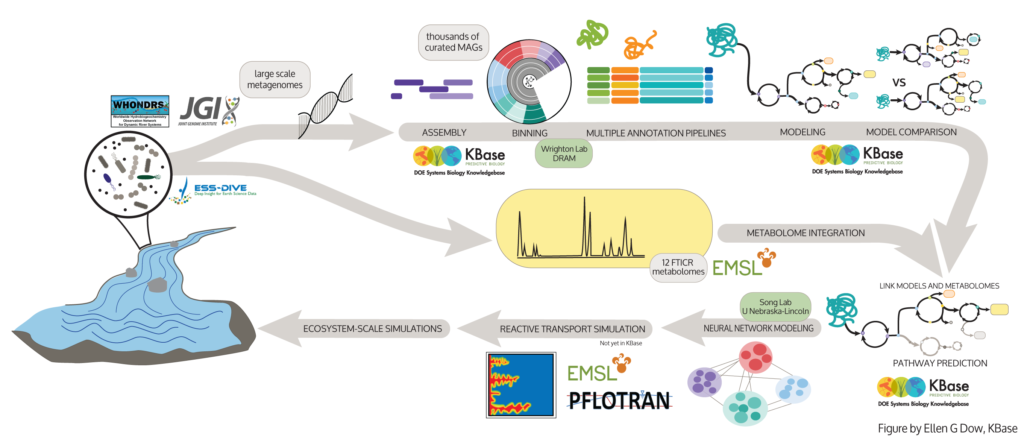
Workflow diagram of Multiscale Microbial Dynamics Modeling from samples through layers of modeling to environment-scale simulations. (Figure by Ellen Dow, KBase)

Course Elements
Welcome to the Multiscale Microbial Dynamics Modeling course. Organizers Drs. Tim Scheibe and Nancy Hess introduce the goals and present an overview of the course modules.
1.1 Introduction
Tim Scheibe & Nancy Hess PNNL
Introduction to the WHONDRS consortium and background information on the data used during the course. Learning concepts include an introduction to metagenomics and applied metagenomics research, with a demonstration of the KBase platform used for many of the analyses.
1.2 WHONDRS Introduction
Amy Goldman & James Stegen PNNL
1.3 Metagenomics 101
Kelly Wrighton Colorado State University
video | presentation | supplementary readings | example hands-on Narrative
1.4 KBase Orientation
Elisha Wood-Charlson LBNL
video | presentation | demo Narrative
1.5 Inferring Biogeochemical Function from Genomes (with DRAM tutorial)
Kayla Borton & Mike Shaffer Colorado State University
video | presentation | supplementary readings | example hands-on Narrative
1.7 Connecting Genes to Function: Trait-based approach
Eoin Brodie, Ulas Karaoz, & Gianna Marshmann LBNL
video | presentation | supplementary readings
1.8 Shared microbial functional traits underpin transferable hydrobiogeochemical processes in river corridors
Mike Shaffer et al. Colorado State University
presentation (presented at AGU 2020)
Exploratory dive into the metabolomics of rivers. This section covers analysis pipelines and approaches to researching metabolomics, as the metabolic inputs and outputs from bacteria, and instruments that identify metabolites.
2.1 Metabolomics in River Corridors
James Stegen & Vanessa Garayburu-Caruso PNNL
video | presentation | supplementary readings
2.2 Metabolomics Analysis Pipeline
Bob Danczak PNNL
video | presentation | supplementary readings | ICR Tutorial (GitHub)
2.3 Connecting Metabolites to Function
Hyun-Seob Song University of Nebraska, Lincoln
video | presentation | supplementary readings | Narrative (DOI: 10.25982/65526.69/1755438) | example hands-on Narrative
2.4 High-resolution Mass Spectrometry (EMSL FTICR-MS Lab)
Will Kew PNNL
Applied section that demonstrates how to create models. The instructors explore the complexities of modeling microbial communities and how to develop metabolic models using KBase.
3.1 Metabolic Modeling Landscape
Hyun-Seob Song University of Nebraska, Lincoln
video | presentation | supplementary readings
3.2 FluxOmics Tools for Metabolic Modeling
Mark Borkum PNNL
video | presentation | supplementary readings
3.3 Building and Using Metabolic Models in KBase
Janaka Edirisinghe ANL
video | presentation part 1 | presentation part 2 | Narrative | example hands-on Narrative
3.4 Integrating Metabolomic Data in Metabolic Models
Chris Henry ANL
3.5 Metabolic Network Modeling and Metabolomics Integration for Comparative Analysis of Biogeochemical Reactions in Multiple River Systems
Aimee Kessel, et al. University of Nebraska, Lincoln
presentation (presented at AGU 2020)
Additional context describing how to scale models from communities to ecosystems using reactive transport models (RTMs) to model environment dynamics. Tutorials explain how to develop RTMs and tie together the concepts of metagenomics and metabolomics for current applications to use RTMs in different riverine systems.
4.1 Incorporating Microbial Processes in RTMs
Tim Scheibe PNNL
4.2 Reactive Transport Modeling Tutorial #1
Kewei Chen PNNL
4.3 ML/AI Methods for Metabolic Modeling in RTMs
Hyun-Seob Song University of Nebraska, Lincoln
video | presentation | supplementary readings
4.4 Reactive Transport Modeling Tutorial #2
Roelof Versteeg & Rebecca Rubenstein Subsurface Insights
4.5 Using R for Hydrologic Data
Michelle Newcomer LBNL
4.6 Omics informed RTM in Marine and Lacustrine System
Christof Meile University of Georgia
4.7 Omics informed RTM in Watershed System
Xingyuan Chen PNNL
video | presentation | supplementary readings
4.8 Omics informed RTM in Wetland Systems
Pamela Weisenhorn ANL
4.9 Opportunities for Science Advancements using WHONDRS in RTM
Tim Scheibe and James Stegen PNNL
The final section covers U.S. DOE user research facilities and their available resources.
Biological and Environmental Research (BER) Facilities and Resources Overview
Joint Genome Institute (JGI)
Rekha Seshadri & Rex Malmstrom
video | presentation part 1 | presentation part 2 | presentation part 3
Environmental Molecular Sciences Laboratory (EMSL)
Will Kew PNNL
Sponsors
 | EMSL, the Environmental Molecular Sciences Laboratory |
|
 | PNNL, Pacific Northwest National Laboratory | |
| DOE, US Department of Energy | ||
 | JGI, Joint Genome Institute | |
| KBase, DOE Systems Biology Knowledgebase | ||
 |  | Microbial Ecosystems Lab at Colorado State University |
| Subsurface Insights | ||
 | WHONDRS, Worldwide Hydrobiogeochemistry Observation Network for Dynamic River Systems | |
 | SBR, Subsurface Biogeochemical Research | |
| ESS-DIVE, Deep Insight for Earth Science Data | ||