2025 Performance Management Metrics End of Year Summary
Read the Full 2025 Summary ReportSummary
The Department of Energy’s Systems Biology Knowledgebase (KBase) was conceived as an open-source, collaborative platform for integrating data, tools, and models to enable researchers to make predictions from complex, multiscale biological data and then share their data, analyses, and insights with others (1). To this end, KBase lowers the barriers to high-performance analysis, supports transparent collaboration, and makes research products citable, reusable, and trackable.
Currently, KBase serves >48,800 users* working in >121,000 KBase Narratives (open, collaborative Jupyter notebook-like analysis workflows), with access to >23 TB of public data and hundreds of tools that operate on diverse biological data types. With that foundation in place, KBase is now addressing its long-term goal of enabling scientists to leverage each other’s work for predictive microbial ecology, thereby allowing greater understanding and manipulation of plant and microbial genomes as a basis for biofuels development and predictive knowledge of carbon and nutrient cycling in the environment. When KBase launched, the community had just begun to grapple with increased rates of production and diversity of data, the growing challenges of doing domain-specific data interoperability at scale, and seeing early hints as to the role machine learning (ML) and artificial intelligence (AI) might play in aiding analyses.
In this year-end summary, we cover KBase’s progress in these areas, with the following details in each of the quarterly Performance Management Metric (PMM) reports:
In FY25, KBase also maintained the platform’s excellent record for up-time of 99% (not including scheduled maintenance windows), user support through our Help Desk, and community engagement through workshops, webinars, and our KBase Educators program (2). KBase also continues to support several User Working Group science research programs, contributes to community-developed biological standards, and streamlines direct transfer of data sets requested by users for analysis in KBase.
*At time of drafting the 2025 PMM, today KBase serves over 55,000 users
Advancing Data Interoperability within KBase
KBase began the FY25 PMM reports by tackling one of the most pressing challenges in systems biology: making heterogeneous biological data interoperable at scale. While the FAIR data principles (Findable, Accessible, Interoperable, Reusable, (3)) have improved discoverability and access, our attempts at data integration and interoperability have revealed that FAIR is necessary but not sufficient for biological data to be truly interoperable. Key FY25 advancements to support better integration and interoperability of biological data include:
● KBase Central Data Model (CDM): An update to KBase’s extensible, object-oriented data model to improve the representation and scalability of genomes, proteins, reactions, taxonomy, and environmental metadata, and make it compatible with a modernized data architecture; we ensure that all datatypes are consistent with respect to each other in how they are referenced and defined.
● BER Data Lakehouse (originally the KBase Data Lakehouse): A more modern and hybrid system that combines the scalability of a data lake with the governance of a warehouse, supports the KBase CDM and other BER tenants (additional details later);
● Pangenome computation: Links tens of thousands of genomes into functional clusters, providing scalable representations for predictive modeling. Instead of one million genomes and one billion genes, the CDM focuses on ~100,000 pangenomes with ~100 million protein families—offering efficiency and comparability across projects.
● Protein Language Models (PLMs): Creation of a PLM-based index to rapidly integrate novel protein sequences into the CDM, enabling the discovery of new genes and functions across DOE datasets.
These developments allow KBase to act as a testbed for BER data science, capable of interoperating with DOE and external data repositories while preparing for AI/ML-driven discovery.
Enabling Team Science at Scale
In the Q2 PMM report, KBase highlighted its role in supporting collaborative, team-oriented science, a key objective of BER. By combining data interoperability with sociological and technical mechanisms for credit, reproducibility, and transparency, KBase seeks to facilitate open, multi-institutional collaboration. Progress in FY25 includes linking samples and data across platforms, continuing to expand data and tool sharing and publishing, and training the next generation of researchers in collaborative, team-oriented science.
Connecting samples and data across platforms
As KBase is not a data generation platform, users have to bring data in from other sources. Case in point, as a BER user facility, the Joint Genome Institute (JGI) generates free high-throughput sequencing, synthesis, and metabolomics data for the community. With a new KBase Data Transfer Service (DTS) integration, users can log into JGI’s Integrated Microbial 2 2025 Performance Management Metrics End of Year Summary Genomes (IMG) portal, search for isolate genomes sequenced by the JGI, and have them directly transferred into their KBase account. Cross-platform user identification leverages ORCID as the authentication ID, which requires the users to have an ORCID associated with both their IMG and KBase user accounts.
One important feature, supporting both team science and open science, is that the DTS system also transfers the “credit metadata,” which includes not only details required for data citation (4) but also information on where the data was generated and who funded it. To illustrate, when a Narrative is prepared for publication, public data not belonging to the Narrative authors that has accompanying credit metadata will be cited by the Narrative DOI. This ensures credit is not missed. KBase also cites all tools used in the workflow that have associated DOIs (typically a software announcement publication). Hence, all who contribute, whether directly or indirectly, are listed.
Another feature often requested is the transfer of sample metadata, to accompany the data file itself. DTS is working with JGI’s IMG and Genomes Online Database (GOLD) and the National Microbiome Data Collaborative (NMDC), which collects and hosts standardized sample metadata for environmental microbiome samples, to streamline the transfer of sample metadata to KBase.
Expanding data and tool sharing and publishing
Biological systems require a lot of data to understand. Reference data provides a consistent, standardized baseline that can be built upon quickly. As such, KBase provides a centralized location for many key reference data sources, including microbial, fungal, and plant genomes from the National Center for Biotechnology Information (NCBI) and several JGI data portals, references for various culture media formulations, and pre-loaded community ontologies that provide standardized labels that enable data interoperability. In FY25, KBase worked to transform our reference data resources into biological relationships, with ontological meaning, into the new KBase CDM and loaded into the BER Data Lakehouse. Testing and exploration are currently underway by members of the KBase, JGI, NMDC, EMSL, and ESS-DIVE teams.
KBase supports ~260 public and private Organizations, providing central hubs for research teams to share Narratives, data, and workflows. KBase Organizations are used by several major BER Scientific Focus Areas (ENIGMA, PMI) as well as university courses and industry partners. A notable academic KBase Organization that exemplifies team science is the GROWdb – the Genome-Resolved Open Watershed database – which hosts >160 river microbiome samples collected by the research community from 90% of U.S. watersheds (5). The GROWdb Org currently has 86 members, including representation from Africa and Central / South America. To support several international workshops, the GROWdb was augmented with samples and data from the Congo River for a workshop at the International Society for Microbial Ecology (ISME, https://www.kbase.us/isme19-workshop) and Lake Yojoa in Honduras for a workshop at an ISME regional meeting in Mexico (https://www.kbase.us/ismelat2025-workshop). GROWdb’s team science approach, which uses KBase for sharing and publishing sample metadata, data, 3 2025 Performance Management Metrics End of Year Summary and Narrative analysis, has inspired several other groups to create similar collections, including a database of produced water microbes (6) and a fermented foods database of ~4,300 microbial species, representing 13,850 genomes clustered at 99% average nucleotide identity (ANI) (7).
Training the next generation of researchers in collaborative, team-oriented science
The KBase Narrative offers a platform for educators to develop workflows within their teaching materials, so that entire classes can reproduce a scientific analysis notebook without requiring coding skills. Since its inception at the start of the COVID-19 pandemic, the KBase Educators program has grown into a robust global community, with a KBase Organization of ~280 members. Working with the community, the Educators program has evolved beyond data analysis in KBase and now provides training for all stages of the scientific process: forming a research question and hypothesis development, experimental design and sample collection, sample processing, and then data analysis and publishing with KBase. The full program, called the Microbiomes In Computational Research Opportunities Network (MICROnet) (https://www.kbase.us/engage/microbiome-training), is funded by the National Science Foundation (NSF), with funds going to train and support educators to purchase supplies and sequencing. Finally, KBase has partnered with the American Society for Microbiology’s Microbiology Resource Announcements (MRA) to highlight student-led research data publications coming out of these programs: https://journals.asm.org/journal/mra/kbase.
Together, these advances reinforced KBase as a trusted environment for collaborative, reproducible, team-driven science.
Building Partnerships and Expanding Resources
In the Q3 PMM report, KBase focused on engaging external partners to expand user-facing capabilities, with highlights spanning data generators, repositories, tool developers, and the KBase User Working Group research community.
Data generators, repositories, and tool developers
KBase has an ongoing partnership with JGI, and many successful examples of co-developed resources and data sharing to enhance KBase’s capabilities. As mentioned above, the Data Transfer Service (DTS) provides a direct transfer link for JGI data into a user’s KBase Narrative. Since its beta launch in Spring 2025, DTS has transferred ~30TB of data, complete with provenance and credit. KBase also makes available ~160 JGI plant genomes (Phytozome), ~180 JGI fungal genomes (MycoCosm), and >230,000 NCBI genomes, as reference databases for users. KBase also has a strong partnership with the Protein Data Bank (PDB). With the successful launch of the joint “Using KBase to access PDB Structures and Computed Structure Models” in late 2022 (available on the PDB YouTube, which has 90,600 subscribers), KBase has continued to improve the representation of protein structures on our platform. Tools are available to map genomes to structures (Import ProteinStructures from a Metadata File in Staging Area), import structures from files (Import Proteinstructures from a Metadata File in Staging Area), and query the RCSB databases for protein structures (Query RCSB databases for protein structures). KBase’s most recent, high-impact tool collaborations have been:
1. Genome Taxonomy Database (GTDB) and their toolkit (8), which enables users to assign unknown genomes and metagenome-assembled genomes (MAGs) to a GTDB taxonomic label. The GTDB Classify App has been run >29,000 times in KBase.
2. Distilled and Refined Annotation of Metabolism (DRAM), which identifies structural and functional elements in DNA sequences of bacteria and archaea, or viruses (DRAM-v), and attaches biological information to those elements (9). The DRAM suite of Apps in KBase have been run >26,400 times.
3. Constraint-based reconstruction and analysis (COBRA) and the associated modeling community have integrated MEMOTE (model quality testing) and ESCHER (model visualization) into KBase, improving the reliability and usability of metabolic models. KBase also adapted our broader metabolic modeling tool suite to use COBRApy, adding new capabilities and enhancing our interoperability with community tools.
Research collaborations: microbiomes, functional genomics, and modeling
Through collaborations with our KBase User Working Groups (UWGs), KBase is able to make significant contributions across a range of research topics. Related to the tools highlighted above, we continue to focus our efforts in three main areas: understanding microbiomes (who is where), functional genomics (what they are likely doing), and modeling (how they interact with each other and their environment). KBase’s most recent, high-impact research collaborations have been:
1. Ecosystems and Networks Integrated with Genes and Molecular Assemblies (ENIGMA) SFA: A BER Science Focus Area (SFA), ENIGMA and KBase regularly partner to develop best practices for sharing data and training early career researchers to use novel technologies. Recent activities have focused on co-developing long-read assembly and strain-level variation tools and tutorials on using those tools in KBase. As an example of their utility, the long-read tool suite in KBase, which includes Apps for read quality and assembly, has been collectively run over 3000 times by over 500 users. We also applied KBase tools to aid in community modeling analysis of nitrate cycling among ENIGMA microbes (10).
2. Microbes Persist SFA: A multi-institutional SFA with leads at LLNL and Northern Arizona University collaborating with KBase, Purdue, JGI, NMDC, and the Genomic Standards Consortium (GSC) to develop community-driven standards for samples from stable isotope probing (SIP) experiments (11).
3. Phenotypic Response of the Soil Microbiome to Environmental Perturbations (Soil Microbiome) SFA: Working closely with the PNNL Soil Microbiome SFA, KBase has incorporated their Omics-enabled global gapfilling (OMEGGA) tool for reconciling genome-scale metabolic models against known growth phenotypes for microbes. By integrating multi-omics data, the tool is capable of gapfilling for correct functions, and associating phenotypes to potential gene candidates. In KBase, the OMEGGA tool has been integrated into the ModelSEED2 release, with the ability to build and improve metabolic models. These Apps, released in 2024, have been run >30,000 times!
4. µBiospheres SFA: KBase has been collaborating with the LLNL µBiospheres SFA for many years to develop tools that build and analyze metabolic models as ensembles, integrating predictions from multiple annotation algorithms. By considering alternative possible annotations for individual genes, we are better equipped to identify areas of agreement or discrepancy, depending on the modeling tool and its underlying database. This analysis capability is highly synergistic with OMEGGA, as the tools can be used together to identify combinations of annotations that lead to an improved fit to experimental data.
5. Persistence Control of Engineered Functions in Complex Soil Microbiomes (PerCon) SFA: The PNNL PerCon SFA has leveraged KBase’s modeling capabilities above to integrate SNEKMER (12), as a protein family fingerprinting tool that annotates against multiple ontologies, including PFAM and PANTHER. The goal of SNEKMER is to extend protein annotation beyond the ∼50–60% of genes that are currently assigned functional annotation with standard annotation techniques.
6. Finally, KBase has collaborated with several other university partners to integrate additional tools for protein annotation, including: GLM4EC, a protein language model-based annotator developed by the Miller lab at University of Colorado, Denver (13); Transyt, a transporter annotator developed by University of Minho (14); and DRAM, an annotation algorithm focused on functions related to global nutrient cycles maintained by the Wrighton lab at CSU (9).
KBase Data Lakehouse: Cross-BER platform integration and guiding principles
The KBase Data Lakehouse architecture offers a strong foundation to support collaborative research at scale. As a working implementation, it also establishes guiding principles, technical patterns, governance practices, and user workflows that can be generalized and adopted by a future BER-wide Data Lakehouse. The approach allows KBase to address the technical, sociological, and cultural barriers to data interoperability, data integration, and team-oriented science while providing a template for other BER programs.
From a technical standpoint, the KBase Data Lakehouse provides a unified data environment that enables researchers to store and access diverse data types in one consistent and scalable system under a cross-functional, common data governance framework. This reduces duplicative pipelines and brittle data hand-offs across disparate systems that often lead to versioning issues and silos. By adopting open data format standards, the platform ensures interoperability across a wide range of analytical tools and programming languages, accommodating the diverse technical backgrounds and preferences within research teams. These design choices form baseline principles for a BER-wide implementation.
The lakehouse architecture incorporates metadata management, schema versioning, and data lineage. These features ensure that data is discoverable, understandable, and adaptable over 6 time, which are critical for a sustained cross-institutional collaboration. Researchers have clearer paths to find and interpret datasets, while data stewards and administrators can maintain oversight and governance without obstructing access or innovation. As a reference implementation, KBase demonstrates how these capabilities can be standardized and scaled across BER.
Beyond the technical capabilities, the KBase Data Lakehouse also addresses sociological and cultural challenges that hinder effective collaboration in data-driven science. A key strength is treating data as a shared and governed resource and making AI-readiness a fundamental goal for harmonization and curation. Centralization, fine-grained access control, and collaborative tools such as AI agents, query editors, and interactive notebooks support a culture of transparency, accountability, and shared ownership. This lowers barriers for researchers who are not data engineers, while empowering advanced users to derive insights and build models without constraints. These practices serve as operating guidelines for a BER-wide lakehouse.
Looking ahead, the KBase Data Lakehouse will be enhanced by the integration of domain-aware AI agents, bringing intelligent, automated support to scientific research. Trained on domain-specific knowledge, these agents can act as co-scientists exploring datasets, generating hypotheses, suggesting relevant literature or collaborators, and orchestrating complex, cross-domain workflows. For example, an agent could analyze metagenomic datasets, link findings to environmental conditions, and recommend experimental directions, shortening the path from question to insight. Embedding these agents directly into the lakehouse demonstrates a path that BER can replicate to democratize advanced analytics and expand interdisciplinary discovery.
Beyond technical innovation, the lakehouse serves as a catalyst for cultural transformation within DOE science. By combining modern governance, collaborative tooling, AI-powered assistance, and transparent, reproducible workflows, it reframes data from isolated products to collective assets. This fosters a team-science mindset where researchers, data engineers, and program administrators collaborate seamlessly, breaking down historical barriers and enabling sustained, scalable cooperation. As a living exemplar, KBase provides the practices and guardrails a BER Data Lakehouse can adopt at the program scale.
As the platform evolves, it will continue to expand in scope and capability. Future enhancements will support additional DOE programs, integrate with national cyberinfrastructure, and bring advanced modeling and simulation environments directly into the lakehouse ecosystem. Governance frameworks will mature to meet the growing complexity of cross-institutional research, while AI-driven prediction and decision-support tools will enable breakthroughs in areas such as climate modeling, bioenergy production, and environmental resilience. These evolutions in KBase are intended to inform and guide the BER Data Lakehouse roadmap.
In essence, the KBase Data Lakehouse is more than a modern data architecture; it is a collaboration enabler and an engine of scientific innovation. By uniting DOE programs under shared principles proven in KBase, it provides the technical and cultural blueprint needed to tackle complex research challenges. With the planned integration of domain-aware AI agents, the lakehouse becomes a living, intelligent ecosystem where data, computation, and collaboration converge, positioning BER to lead the future of open, team-oriented, AI-powered science and accelerating the path from data to discovery.
KBase Data Lakehouse: high-level milestones
KBase’s Data Lakehouse has progressed from a central data model idea to a running platform that teams are already using. More than a KBase deliverable, it establishes guiding principles, including governance, open formats, AI-readiness, agentic workflows, and user-centric UX that a BER-wide Data Lakehouse can adopt.
KBase subject-matter experts are actively building the Central Data Model on the Lakehouse harmonizing schemas, metadata, and provenance to enable reliable cross-study analysis. In parallel, the KBase AI team is delivering future-ready agentic capabilities (e.g., MCP-backed services, multi-agent orchestration, and reasoning-trace/lineage retention) on this platform for KBase and, ultimately, the broader BER ecosystem. Following this foundation work, we have onboarded SFA minitenants, including ENIGMA and Phage Foundry, along with partner groups such as the ARPA-H team, validating the multi-tenant model and shared governance patterns across programs.
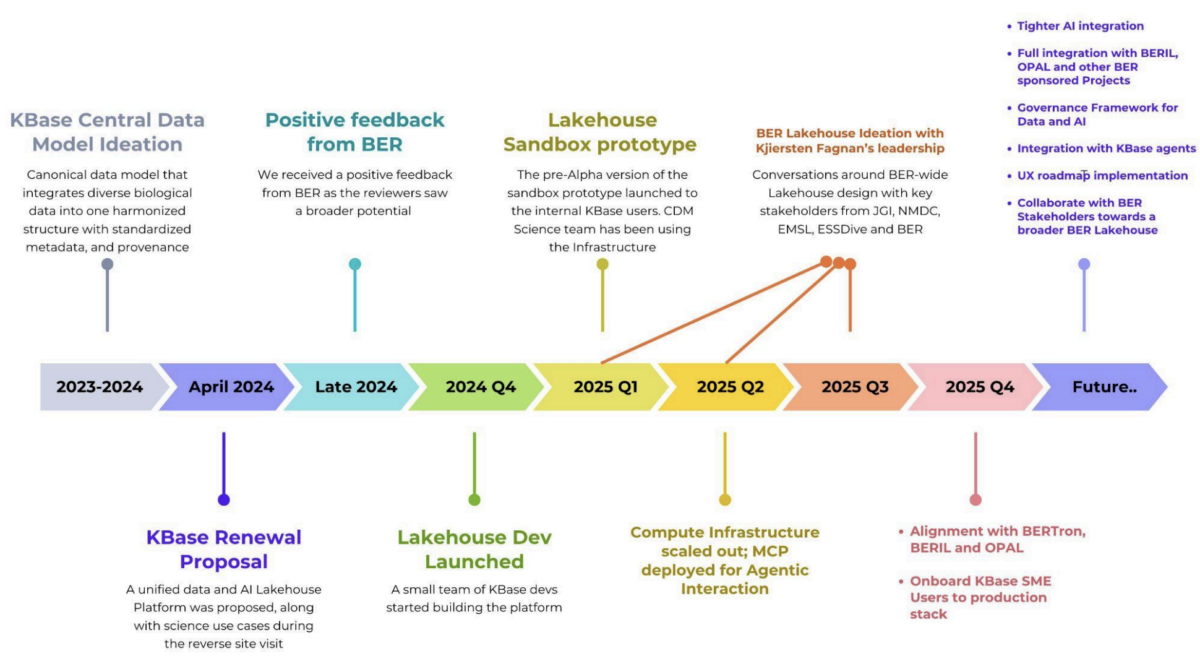
Milestones at a glance: the timeline depicted in the figure above traces the KBase Data Lakehouse journey from early Central Data Model ideation through proposal, build-out, sandbox and query hub launches, APIs and MCP services, and into production readiness. This effectively demonstrates how the KBase team turned vision into a working platform and a reusable blueprint for BER. This KBase Data Lakehouse Journey demonstrates not just how KBase built the lakehouse, but how BER can scale it through adoption of the same principles, architecture patterns, and onboarding approach to accelerate team science across the entire BER ecosystem.
Incorporating AI/ML for Next-Generation Capabilities
The final quarter of FY25 marks the initial incorporation of AI/ML approaches by KBase. Building on interoperability (Q1), collaboration (Q2), and external partnerships (Q3), KBase is now embedding AI/ML into its workflows to accelerate predictive biology. The reader is referred to the Q4 PMM for details.
Key emerging areas include:
● KBase Narrative Agent: Many users new to computational biology often struggle with the lack of familiarity with specific tools and workflows to conduct their analysis. To address this gap, a prototype AI-driven system has been built, which suggests to the users which tools and workflows facilitate their analysis and are likely to answer their scientific questions of interest.
● KBase Gene Function Agent: To overcome the limitations of current gene annotation pipelines, which may have conflicting or incomplete information, this approach integrates multiple lines of evidence and applies AI reasoning to generate improved gene function predictions.
● Generative AI Foundation Model for Genome Composition: A challenge with draft microbial genomes and metagenome-assembled genomes (MAGs) is that they are often incomplete or fragmented, making it difficult to recover accurate functional profiles. KBase is developing a generative AI foundation model for microbial genome composition by training it directly on species-level pangenomes spanning thousands of microbial lineages in an attempt to build a consensus where possible.
● Protein Language Model for Sequence Search: Determining protein similarity has been a key component of numerous genome annotation, comparative genomics, and omics data analysis pipelines. But, this computation has become increasingly expensive to run with the growing volume of available reference genomic data. Protein language models (PLMs) provide a means of making protein similarity search vastly more scalable and more sophisticated, with benefits for accuracy. These approaches are being actively explored as a foundation model asset in the KBase Lakehouse.
● Machine Learning Classifiers for Microbial Phenotype Prediction: To answer the question of whether a microbe can grow on a particular carbon source based on knowledge of its genome has been of long-term interest in biology. KBase has developed ML classifiers that predict microbial phenotypes from genomic features.
These models are trained on an extensive dataset of 819 microbial genomes tested across 242 carbon sources, drawing on multiple experimental resources (ATLeaf, Biolog, Marine, and PMI datasets).
The classifiers integrate diverse genomic annotations, including RAST, KOFAM, UniProt, eggNOG, and UniRef, converting gene-level information into feature matrices for machine learning for improved phenotype prediction. 9 2025 Performance Management Metrics End of Year Summary These efforts lay the foundation for FY26 priorities, where AI/ML will become central to KBase’s role as a next-generation knowledge environment for BER.
FY25: Platform stability, user satisfaction, community engagement
While KBase’s main efforts have been focused on data interoperability and building partnerships to enable team science and AI, KBase also continues to support the current production platform and our user community. Throughout FY25, we have maintained a platform up-time greater than 99% (outside of scheduled downtimes for maintenance). The KBase Help Desk has supported 50 user questions, resolved 43 bug reports, and has an overall satisfaction rating of 4.7 out of 5. KBase also hosted 36 workshops and webinars in FY25, reaching 1400 members of our user community. Training ranged from genome assembly of microbial isolates to metabolic modeling of microbial community dynamics, for users across a range of career stages, including undergraduates and senior researchers at universities and national laboratories. KBase training sessions are made available on the KBase YouTube channel (https://www.youtube.com/DOEKBase), which has ~1500 followers. Figure 1 is an infographic summarizing key platform/user statistics.
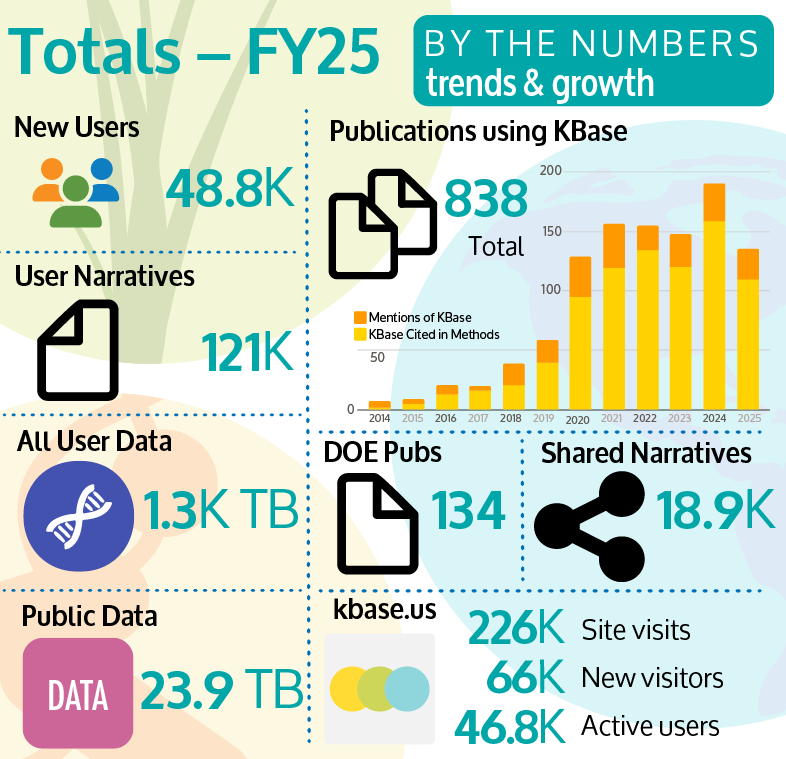
Figure 1. Key KBase metrics
Summary
KBase remains a cornerstone of DOE’s systems biology research ecosystem, ensuring that data and models are not only accessible, but actionable, to advance predictive understanding of plants, microbes, and environmental systems. The KBase community continues to grow and demonstrate the power of community open science through contributions of tools; donation, curation, integration, and sharing of data; and production of analyses, many of which progress to published form in both archival journal format and/or as citable and trackable published KBase Narratives. 10 2025 Performance Management Metrics End of Year Summary This year we have seen a significant deepening of the core genotype-to-phenotype prediction systems, incorporating improved taxonomic classification linked to function through our pangenome efforts and collaboration with GTDB. Expansion and deepening of genome annotation at the gene and trait levels, which are critical for genome-scale modeling. These data and assertions are now driving stand-up and improvement of services for ML-based, knowledge-based, and mechanistic models linking genotype to phenotype in microbes. This work is now forming the basis for testing the other major innovation in this past year: the stand-up of our first lakehouse prototype and agentic layers linking lakehouse to KBase designed to help data query, execute automated improvements of things like gene annotations, and aid users in designing, and executing research on the system. While nascent, this stands as a mode of the future of biological data science across BER programs. The prototype has been of interest to several BER SFAs, BRaVE projects, and even an APRA-H program. Even just as guests on our nascent datalakehouse/AI infrastructure, they have been able to upload, query, and execute demonstrations of this new technology. We are also very gratified and pleased to participate with the standup of the collaborative BER efforts in integrative data science: BERTron, the emerging cross-facility data lakehouse (currently called BERDL), and the pilot AI project BERIL and OPAL. We see an incredible synergy with these efforts that should accelerate science across BER.
References
1) Arkin A, Cottingham R, Henry CS, et al. (2018). KBase: The United States Department of Energy Systems Biology Knowledgebase. Nat Biotechnol, 36, 566–569. https://doi.org/10.1038/nbt.4163
2) Dow EG, Biller SJ, Paustian T, Schirmer A, Sheik CS, Whitham JM, Krebs R, Goller CC, Allen B, Crockett Z, Arkin AP (2021). Bioinformatic teaching resources – for educators, by educators – using KBase, a free, user-friendly, open source platform. Frontiers in Education, 6, 711535. https://doi.org/10.3389/feduc.2021.711535
3) Wilkinson MD, Dumontier M, Aalbersberg IJ, Appleton G, Axton M, Baak A, Blomberg N, Boiten J, Bonino L, Bourne PE, Bouwman J, Brookes AJ, Clark T, Crosas M, Dillo I, Dumon O, Edmunds S, Evelo CT, Finkers R, Mons B (2016). The FAIR guiding principles for scientific data management and stewardship. Scientific Data, 3(1), 1–9. https://doi.org/10.1038/sdata.2016.18
4) ESIP Data Preservation and Stewardship Committee (2019). Data citation guidelines for Earth science data, version 2. ESIP. https://doi.org/10.6084/m9.figshare.8441816.v1
5) Borton MA, McGivern BB, Willi KR, et al. (2025). A functional microbiome catalogue crowdsourced from North American rivers. Nature, 637, 103–112. https://doi.org/10.1038/s41586-024-08240-z
6) Schweitzer HD, Tinker KA, Barnhart EP, Akob DM, Crockett ZR, Gulliver D (2025). Produced water DNA database (PW-DNA): utilizing KBase to generate an environmental-specific curated molecular database. Dataset, Sep 2025. https://doi.org/10.25982/156785.278/2588866
7) McDaniel EA (2025). Fermented foods microbial genomes database. Jun 2025. https://doi.org/10.25982/218406.47/2569606
8) Chaumeil PA, Mussig AJ, Hugenholtz P, Parks DH (2020). GTDB-Tk: a toolkit to classify genomes with the genome taxonomy database. Bioinformatics, 36(6), 1925–1927. https://doi.org/10.1093/bioinformatics/btz848
9) Shaffer M, Borton MA, Bolduc B, Faria JP, Flynn RM, Ghadermazi P, Edirisinghe JN, Wood-Charlson EM, Miller CS, Chan SHJ, Sullivan MB, Henry CS, Wrighton KC (2023). 11 2025 Performance Management Metrics End of Year Summary kb_DRAM: annotation and metabolic profiling of genomes with DRAM in KBase. Bioinformatics, 39(4), btad110. https://doi.org/10.1093/bioinformatics/btad110
10) Carr AV, Otwell AE, Hunt KA, Chen Y, Wilson J, Faria JP, Liu F, Edirisinghe JN, Valenzuela JJ, Turkarslan S, Lui LM, Nielsen TN, Arkin AP, Henry CS, Petzold CJ, Stahl DA, Baliga NS (2025). Emergence and disruption of cooperativity in a denitrifying microbial community. The ISME Journal, 19(1), wraf093. https://doi.org/10.1093/ismejo/wraf093
11) Simpson A, Wood-Charlson EM, Smith M, Koch BJ, Beilsmith K, Kimbrel JA, Kellom M, Hunter CI, Walls RL, Schriml LM, Wilhelm RC (2024). MISIP: a data standard for the reuse and reproducibility of any stable isotope probing-derived nucleic acid sequence and experiment. GigaScience, 13, giae071. https://doi.org/10.1093/gigascience/giae071
12) Chang CH, Nelson WC, Jerger A, Wright AT, Egbert RG, McDermott JE (2023). Snekmer: a scalable pipeline for protein sequence fingerprinting based on amino acid recoding. Bioinformatics Advances, 3(1), vbad005. https://doi.org/10.1093/bioadv/vbad005
13) Davoudi S, Henry CS, Miller CS, Banaei-Kashani F (2025). EC-Bench: a benchmark for enzyme commission number prediction. bioRxiv, 2025.06.25.661207. https://doi.org/10.1101/2025.06.25.661207
14) Cunha E, Lagoa D, Faria JP, Liu F, Henry CS, Dias O (2023). TranSyT: an innovative framework for identifying transport systems. Bioinformatics, 39(8), btad466. https://doi.org/10.1093/bioinformatics/btad466
