Protein Data Bank
KBase – PDB Integration Framework
KBase has partnered with the Research Collaboratory for Structural Bioinformatics (RCSB) Protein Data Bank (PDB, https://www.rcsb.org/) to provide researchers with enhanced connections between structural data and “multi-omics” data, such as genomics, metagenomics, transcriptomics, and metabolomics.
The partnership began with a series of workshops to identify community needs and gather feedback, which guides the prioritization and development of tools and connections between KBase and PDB. The aims of this collaboration are:
- Host a community workshop series
- Establish connection to expose 3D structures to the KBase platform
- Develop KBase tools (Apps) and UI to enable users to interact with PDB data
- Establish connect to expose KBase data to PDB
- Develop web interface in PDB to enable users to explore KBase data
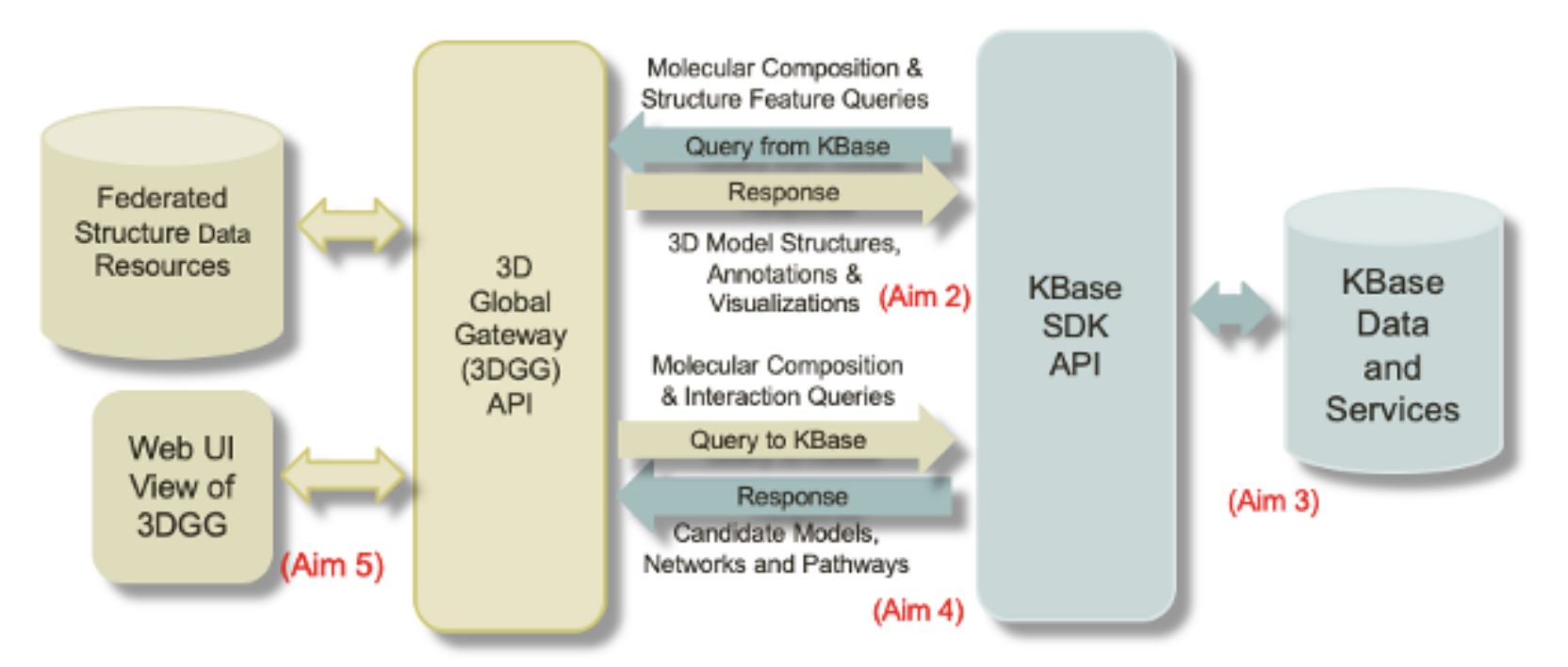
By linking KBase to PDB, structure data will be fully integrated, with realtime updates as new structures are added to PDB, and the protein contact map information will support new synthetic biology workflows. By linking PDB to KBase, users will be able to access the broader systems biology context to proteins of interest, including taxonomic relationships between organisms, co-occurrence networks of proteins and genomes, and associated multi-omics data sets.
Functionality and Tools
- Query RCSB databases for protein structures: Query Research Collaboratory for Structural Bioinformatics (RCSB) databases for a list of protein structures. (Beta-only)
- Import Protein Structures: Import PDB files from upload staging area into a KBase Narrative. (Beta-only)
- Import PDB Metadata into KBase Genome (Beta-only)
Additional Information
Meet the Members
Principal Investigators: Steven K. Burley1, John D. Westbrook1#, Chris Henry2,3
Developer Team: Sam Seaver2,3, Qizhi Zhang2,3, Claudia Lerma-Ortiz2,3
Affiliations: 1Rutgers, The State University of New Jersey, 2Argonne National Lab, 3KBase
#passed away
Resources
- Using KBase to Access Protein Structures and Models (YouTube playlist)