Quarterly User Update (January – March 2026)
This community update covers our 2026 activities and events from January through March – covering announcements and highlights; and celebrating accomplishments from our team, users, and collaborators.
Community Highlights
Our latest stories about the KBase community cover new resources available on the platform to how KBase is used across the globe. This quarter’s highlights include Fulbright Fellow Maureen Morrow (https://www.kbase.us/news/maureen-morrow/) sharing her contributions to training future researchers, Elizabeth McDaniel (https://www.kbase.us/news/elizabeth_mcdaniel/) on the newly-released fermented foods database, and Rian Pierneef (https://www.kbase.us/news/rian-pierneef/) on running champion-driven workshops across Africa that use KBase.
If you are interested in a KBase Highlight on your work or projects, email us at [email protected].
Platform Updates
The first version of the JGI-KBase Data Transfer Service (DTS) can be used to push IMG genomes from JGI to KBase for further analysis. Details and a demonstration on how to use the DTS can be found here: https://www.kbase.us/news/jgi-kbase-dts/.
KBase Publications
The platform and team behind KBase continues to support users to analyze their data and follow the FAIR (findable, accessible, interoperable, reusable) data principles through to publication. Studies published this quarter included:
- Developing a KBase-specific resource for metabolic modeling of neutrophilic iron-oxidizing bacteria (Tothero et al. 2026).
- Characterization of novel Nitrobacter vulgaris strain MLSD-S22 isolated from a nitrate- and heavy-metal-contaminated subsurface environment with a complete genome and plasmids capable of nitrous oxide reduction (Flinkstrom et al. 2026).
- Insights and hypotheses from metagenomic characterization of agricultural microbial inoculant “Jeevamrit” (Jain et al. 2026).
All publications using KBase are available at: https://www.kbase.us/research. If we are missing your publication citing KBase in your methods, please let us know at [email protected].
New KBase Citation!
There is a new publication to cite for KBase! You can now include the following when referencing KBase in your methods:
E.M. Wood-Charlson, C. S. Henry, P. S. Dehal, G. Mahmud, B. H. Allen, K. Beilsmith, D. D. Blair, S. Canon, M. Cashman, D. Chivian, R. Cottingham, Z. Crockett, E. G. Dow, M. Drake, J. N. Edirisinghe, J.P. Faria, A. Freiburger, T. Gu, P. Gupta, A.J. Ireland, S. Jungbluth, R. Kamimura, K. Keller, A. Khan, D. Kishore, D. Klos, F. Liu, D. Lyon, C. Neely, K.L. O’Grady, G. Price, P. Ranjan, W.J. Riehl, B. Sadkhin, S. Seaver, G.A. Terry, Y. Wang, P. Weisenhorn, Z. Yang, S. Yoo, A.P. Arkin. “KBase: Open-source Platform for Collaborative Biological Data Analysis and Publication.” Journal of Molecular Biology (2026) 169676. [doi: 10.1016/j.jmb.2026.169676]
Summary
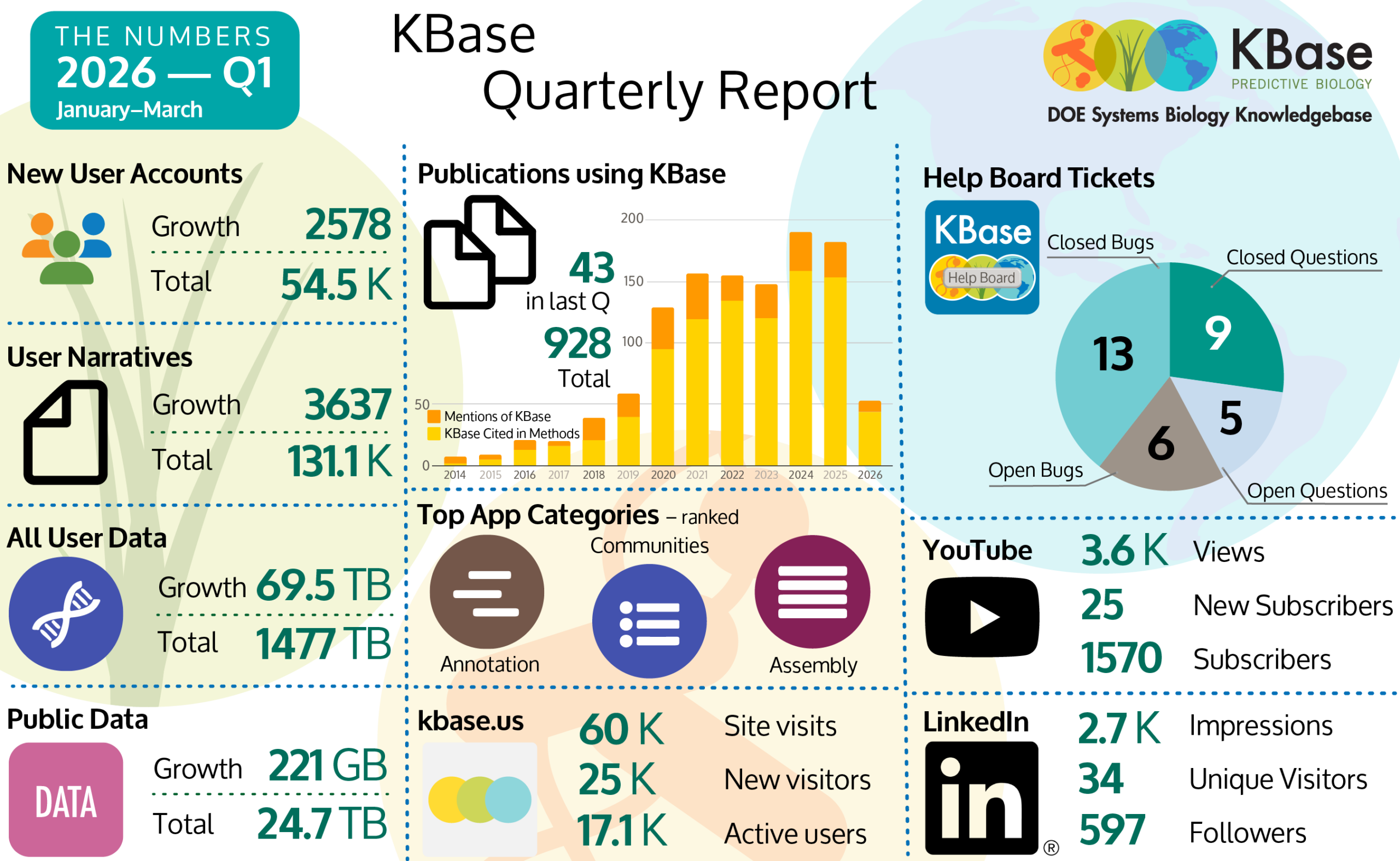
To close out our update, we have our platform 2025 Performance Metrics Report and below is a graphical summary documenting our Q1 January – March 2026 user numbers. Keep doing great science in KBase. Remember, the KBase team is here to support you!