User Working Groups
Partnering with researchers and data scientists, KBase is working to advance a joint mission of sharing data and knowledge with the science community.
Microbiome
Microbes are diverse and pervasive, existing across space and time. Collaborations focused on understanding microbiome communities explore what microbes are where, and what they are doing there. Much of this work delves into the genetic potential of organisms, especially how they are capable of interacting with each other and their surrounding environment. Understanding the potential of microbiomes helps us predict ecosystem resilience and how they might respond to change.
Research within the Ecosystems and Networks Integrated with Genes and Molecular Assemblies (ENIGMA) project, led by Lawrence Berkeley National Lab (LBNL), emphasizes achieving a multiscale, causal, and predictive understanding of microbial biology. This includes understanding the relationship between molecules, microbes, communities, and the impact on the ecosystems they inhabit
To find out more about ENIGMA community developer contributions to KBase, check out the ENIGMA collaboration page.
ENIGMA SFA home page
The Microbes Persist: Systems Biology of the Soil Microbiome (Microbe Persist) Science Focus Area, led by Lawrence Livermore National Lab (LLNL), explores how microbial ecophysiology and population dynamics, as well as biotic-abiotic interactions regulate the microbial resilience soil under changing soil moisture conditions.
To find out more about Microbe Persist community developer contributions to KBase, check out the Microbes Persist collaboration page.
Microbes Persist SFA home pageThe Bacterial-Fungal Interactions Science Focus Area (SFA) at Los Alamos National Lab (LANL) uses a multi-omics approach to address knowledge gaps and gain a predictive understanding of the bacterial-fungal interactions in the soil, with the goal of understanding how microbial interactions affect ecosystem services such as plant productivity, nutrient cycling, and carbon flux.
To find out more about Bacterial-Fungal Interaction community developer contributions to KBase, check out the Bacterial-Fungal Interaction collaboration page.
Bacterial-Fungal Interactions SFA home pageThe Microbial ecoSystems Laboratory, led by Dr. Kelly Wrighton at Colorado State University, focuses on using knowledge about microbiomes to better manage ecosystem function. Through employing cutting edge technology to inventory microbes from a range of environments, the group creates microbiome profiles to understand and predict community function under changing environmental conditions.
To find out more about Microbial ecoSystems Laboratory’s contributions to KBase, check out the Microbial ecoSystems Laboratory collaboration page.

Metabolism
Collaborations around metabolic networks explore how cells (free-living, host-associated, or multicellular) communicate with other cells, collaborate or compete for resources, and interact with their environment. Research in this area connects the genetic potential of cells/communities to environmental conditions and aims to understand how life responds to change.
The µBiospheres Science Focus Area (SFA): Community systems biology of microscale interactions for sustainable bioenergy at Lawrence Livermore National Lab (LLNL) aims to develop a predictive framework around the formation and maintenance of phototroph–microbial interactions. The SFA looks microbial consortia from a system’s resource perspective to better understand sustainable bioenergy cultivation.
To find out more about µBiospheres community developer contributions to KBase, check out the µBiospheres collaboration page.
µBiospheres SFA home page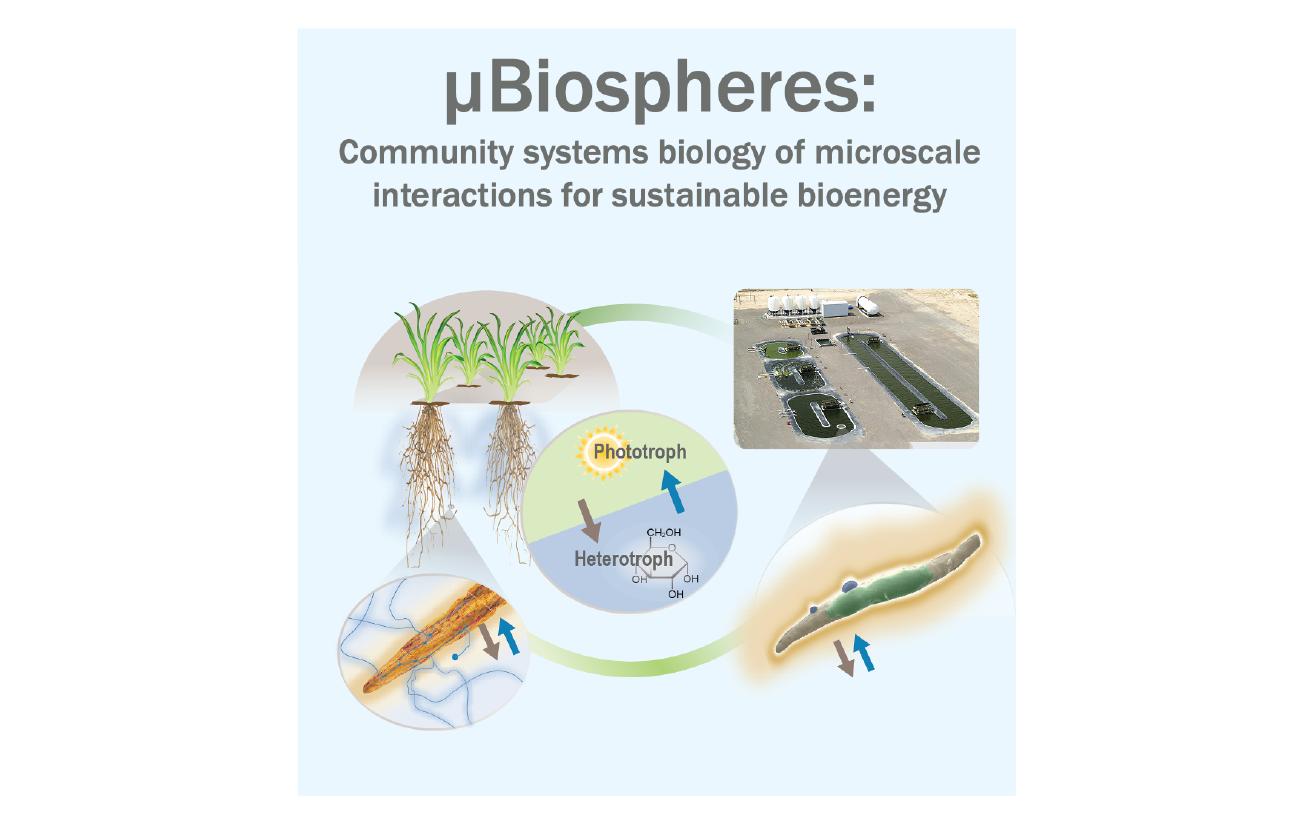
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations (Soil Microbiome) Science Focus Area (SFA), led by Pacific Northwest National Laboratory (PNNL), is tackling the challenge of understanding how soil ecosystems (plants, microbes, nutrients, etc.) respond to changes in the environment, such as drought.
To find out more about the Soil Microbiome community developer contributions to KBase, check out the Soil Microbiome collaboration page.
Soil Microbiome SFA home page
The Persistence Control of Engineered Functions in Complex Soil Microbiomes (PerCon) Science Focus Area (SFA), led by Pacific Northwest National Laboratory (PNNL), explores genome-scale engineering of rhizosphere microbiomes as a way to create resilient and productive bioenergy crops. Understanding the genetics, metabolomics, spatial distribution of microbes in the rhizosphere will help PerCon create and establish persistent host-microbe symbioses.
To find out more about the PerCon community developer contributions to KBase, check out the PerCon collaboration page.
Persistence Control SFA home page
Functional Genomics
Functional genomics collaborations investigate a range of multivariate genotype-phenotype relationships to study how genes or genomic regions are predictive of biological function in a particular environmental context. Associated data include genomic and phenotypic assessments, along with integrations of epigenomic, transcriptomic, proteomic, metabolomic and other data necessary to improve predictions. Studies can be direct or indirect, with the goal of linking function to genes and understanding the impact of genetic variation in populations.
The Biofuels and Protein Structure Science Focus Area (SFA) at Oak Ridge National Laboratory (ORNL) aims to understand how to efficiently develop renewable biofuels from lignocellulosic biomass. This includes understanding the process of biomass deconstruction, the enzymes responsible and the byproducts they generate to maximize biofuel production.
To find out more about Biofuel and Protein Structure community developer contributions, check out the Biofuels and Protein Structure collaboration page.
Biofuels and Protein Stucture SFA home pageThe Plant Microbe Interfaces (PMI) Science Focus Area (SFA) at Oak Ridge National Laboratory (ORNL) explores the dynamic interface between plants, microbes, and their environment. With a focus on Populus (poplar trees), PMI aims to understand carbon sequestration and cycling in terrestrial systems, and how those processes may respond to environmental changes and provide renewable sources of energy.
To find out more about the PMI community developer contributions to KBase, check out the PMI collaboration page.
Plant Microbe Interfaces SFA home page
The Ware lab at Cold Spring Harbor Laboratory (CSHL) looks at functional and comparative plant genomics, with an emphasis on agricultural crops, and develops tools for the plant genomics research community.
To find out more about the Ware lab contributions to KBase, check out the Ware lab collaboration page.
Ware Lab at CSHLA core component to linking genotype to phenotype includes linking genomics to proteins and their structures. The Protein Data Bank (PDB) provides 3D protein structures derived from experimental and computational approaches, and KBase provides diverse systems biology data that provides context to protein function.
To find out more about how PDB and KBase are connecting these complementary resources, check out the PDB collaboration page.
Protein Data Bank
Data Science
Developing detailed models of biological function within an environmental context enables researchers to explore prediction and design of ecosystem services that address challenges of changing conditions, land and water use, and other anthropogenic impacts. Data science employs a diverse and sophisticated set of statistical transformations and machine learning methods built from a suite of measurement technologies, spanning the molecular to the ecological, that are being linked together by increasingly sophisticated experimental designs.
In collaboration with the Daniel Jacobson Lab at Oak Ridge National Lab (ORNL), an exascale network prototype for the model organism Arabidopsis thaliana using layered data that encompasses DNA, RNA, proteins, metabolites, phenotype, and environmental conditions.
To find out more about the Exascale Network community developer contributions to KBase, check out the Exascale Network collaboration page (coming soon).
This activity is still in beta testing, and we are looking for beta testers! Sign up here.
Data science methodology requires well curated and consistent data sources for success. To support the community’s data science research efforts, KBase and the Joint Genome Institute (JGI) are developing shared microservices so our users can expect consistent and reliable results across our platforms.
To find out more about how JGI and KBase are working together, check out the JGI collaboration page.
Joint Genome Institute